Our agentic pipeline uses LLMs to construct rubrics — structured specifications for automatically transforming raw, heterogeneous inputs into powerful representations for efficient downstream supervised learning.
Global Rubric Pipeline
Label-stratified k-means clustering in embedding space selects 40 diverse, representative examples (20 per class) as medoids from the training set.
An LLM agent analyzes the 40 medoid EHRs in-context and synthesizes a task-specific rubric — a structured template defining what evidence to extract.
The output is a systematic rubric ℛ with sections like Demographics, CV Risk Factors, Comorbidities, Temporal Trends, and Alert Flags.
An LLM fills in every rubric field for each patient using only data from their EHR. High fidelity, but requires one LLM call per example.
An LLM generates a deterministic Python parser that applies the rubric via string/regex matching — no LLM calls needed at inference time.
An LLM generates a script converting rubric outputs into numeric feature vectors, enabling standard ML models like XGBoost.
Full prompt for rubric synthesis (Panel B)
You are a medical expert designing a structured rubric for a clinical prediction task.
## Task
- Name: {task_name}
- Query: {task_query}
## Context
You will be given {40} labeled patient EHR examples ({20} positive, {20} negative). Another model will later use your rubric to transform new patient EHRs into structured summaries, which will then serve as input to a supervised classifier.
## What You Must Do
Study the examples below. Combine what you observe in them with your medical knowledge to design a rubric template — a set of named fields that, when filled in for any patient, produce a structured summary optimized for this prediction task.
The rubric should:
- Be data-driven and discriminative. Identify which features, patterns, and interactions actually separate the positive and negative cases.
- Be structured and consistent. Every rubricified output must follow the same field names and order.
- Extract facts only. The evaluator filling in the rubric must NOT make predictions, assign risk levels, or draw conclusions.
- Be concise. Focus on extracting information relevant to the task, not reproducing the entire EHR.
## Examples
{Positive and negative EHR serializations}
## Output
Output ONLY the rubric template itself — the instructions another model will follow to transform a patient EHR. No preamble, no explanation. The template must be self-contained and directly usable.
Two Types of Rubrics
Global Rubrics
A single shared rubric is synthesized from a small subset of examples and applied uniformly to all inputs.
- Fixed schema — easy to audit and reproduce
- Can be applied via deterministic parser scripts (zero LLM cost at inference)
- Convertible to tabular features for standard ML
Local Rubrics
A task-conditioned summary is generated per example by the LLM, producing structured sections like Patient Snapshot, Risk Factors, and Protective Factors.
- More flexible, potentially higher fidelity
- Captures patient-specific nuances
- Requires an LLM call per example
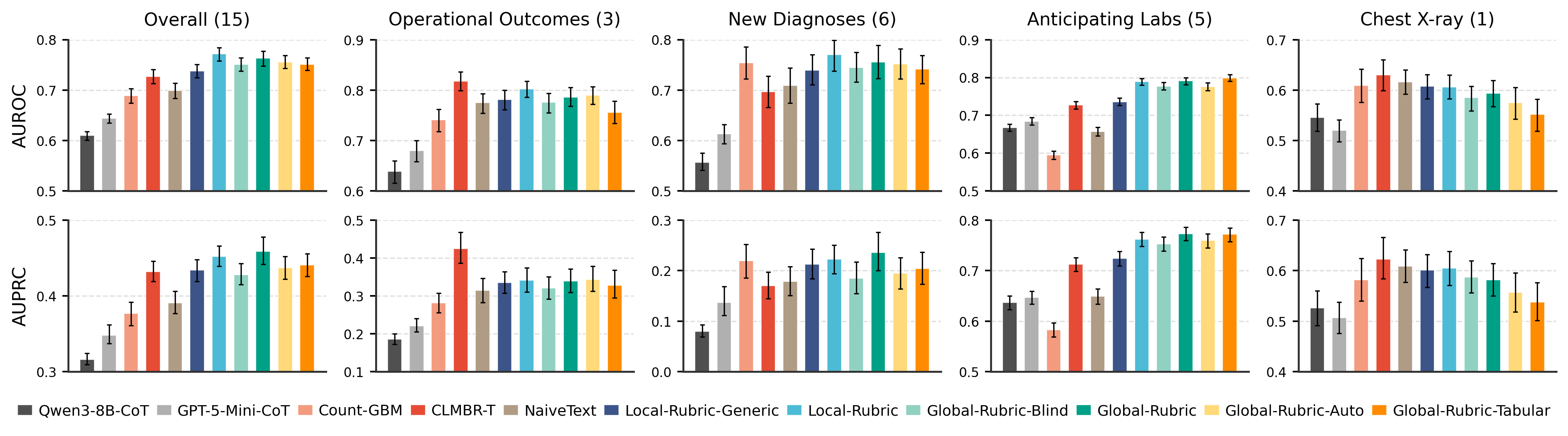
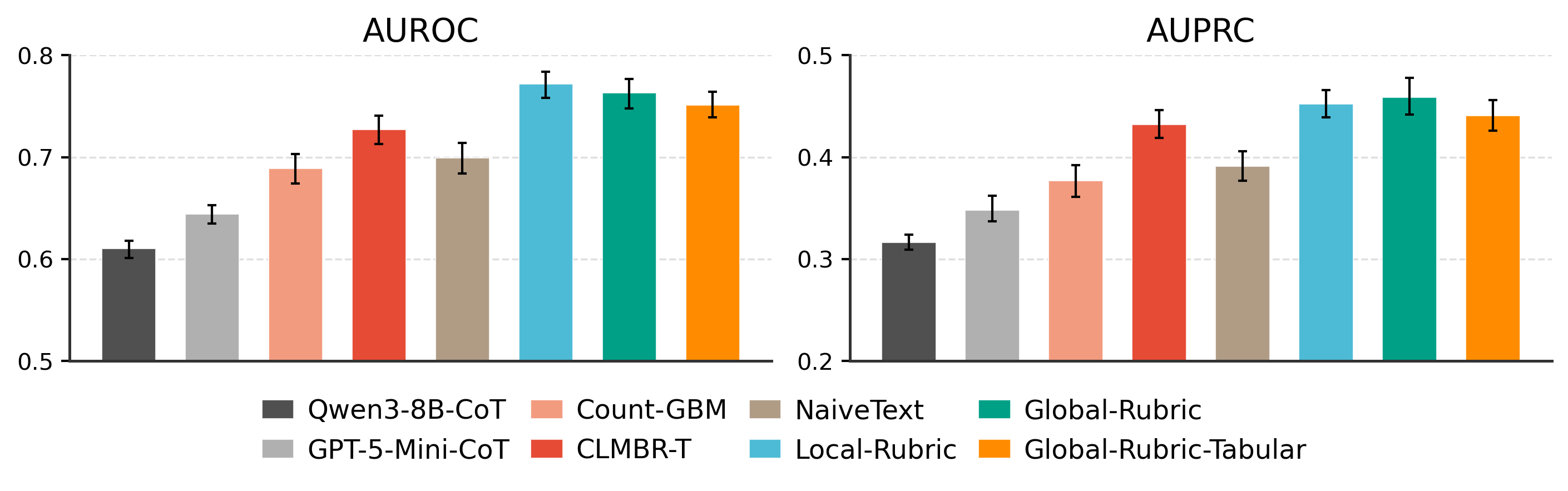
Representation Comparison
The rubric transforms a raw, noisy text serialization into a structured, evidence-organized representation:
## Patient Demographics
- Patient age: 78, FEMALE [...]
### Inpatient Visit (14 days to pred. time)
Conditions: Acute posthemorrhagic anemia [...]
Medications: furosemide 20 MG [...]
### ER Visit (87 days before)
Conditions: Benign essential hypertension [...]
Medications: ondansetron, nitroglycerin [...]
1. Patient Snapshot
27 yo hispanic male. Recurrent cardiology visits [...]
2. Main Risk Factors
- Congenital coronary artery anomaly [...]
- Tobacco exposure (smokeless) [...]
3. Protective Factors
- Young age (27), Normal BMI (21-22) [...]
6. Overall Risk Impression
Elevated risk of acute MI [...]
§3. Demographics
55 | FEMALE | [...]
§6. Recent Cardiac Symptoms (last 365d)
- Chest pain/angina: No
- Dyspnea: Yes [...]
§12. Other Relevant Labs
- Creatinine: 1.12 (2023-12-02)
- eGFR: No data [...]
§17. Known Risk Factors
- Diabetes: No, Hyperlipidemia: Yes
- Family hx of premature CAD: Unknown [...]